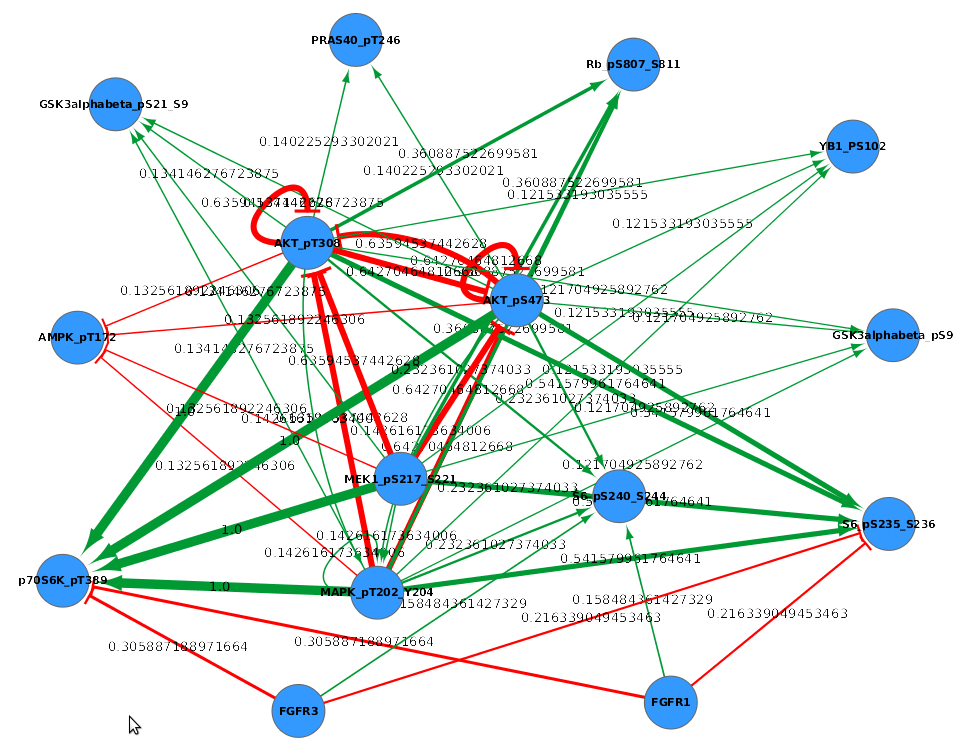
Actor A appears in a film with Actor B.Other kinds of relationships you could describe in an edge list: All that matters here is that each row in my spreadsheet describes a connection between two entities. The columns could be reordered so that the Films column precedes the Actors column it doesn’t matter. In the image below, my edge list contains two kinds of things: films and actors. The objects in the first column should be connected in some way to the objects in the second column. At its most basic, Cytoscape wants a spreadsheet containing two columns. The most important thing to understand about Cytoscape (or, really, most network analysis tools) is the format in which you need to structure your data. (You should also, however, be able to follow these tutorials if you have your own data you’d like to use.) 1. It contains actors and films from the silent era of American race film ( source). We’ll also look at how to transform a bimodal graph into a unimodal graph, and we’ll walk through the process of publishing your graph to the web.įor this tutorial, please download this spreadsheet and save it somewhere you can find it again. We’ll start by building a basic graph and then work with more advanced features, like attributes and calculations. Along the way, you’ll come to understand some of these network concepts more fully. This set of tutorials will walk you through the process of building a network graph in Cytoscape. In addition, Scott Weingart has a great introduction to network analysis. Cytoscape is not too hard to use, but it won’t make much sense unless you have a sense of some basic network analysis vocabulary and concepts. Get a unimodal network from a bimodal networkĬytoscape is a tool for viewing and analyzing networks (meaning, in this case, any group of entities that are connected in some way). 
Tableau 2: Basemaps, data layers, and geolocation.Messing around with the Topic Modeling Tool.Pro-tip: depending on your data, it may be better to balance your color range over negative and positive values bounded by the largest min or max data value, so that color intensity scales similarly in both directions. setNodeColorMapping('an', data.values, lors, style.name=style.name) lors <- c(rev(brewer.pal(length(data.values), "RdBu")))įinally, we use the handy mapVisualProperty function to construct the data object that CyREST needs to specify style mappings and then we’ll send them off to Cytoscape with updateStyleMappings. (length(data.values), colorblindFriendly=TRUE, type="div") # div,qual,seq,all Next, we use the RColorBrewer package to help us pick good colors to pair with our data values. The first step is to grab the column data from Cytoscape and pull out the min and max to define our data mapping range of values. Now let’s update the style with a mapping for mean expression. Pro-tip: don’t forget to setCurrentNetwork() to the correct parent network before getting table column data and making selections. setCurrentNetwork(network="STRING network - ovarian cancer")ĬommandsPOST('diffusion diffuse') # diffusion!ĬreateSubnetwork('selected',subnetwork.name = 'top disease diffusion') …from diffusion algorithm starting with top 3 genes, using the network connectivity in a more subtle way than just first-degree neighbors. Top.nodes <- row.names()),3)]ĬreateSubnetwork('selected', subnetwork.name='top disease neighbors') # selected nodes, all connecting edges (default) setCurrentNetwork(network="STRING network - ovarian cancer") …from first neighbors of top 3 genes, using the network connectivity together with the data to direct discovery. …of connected nodes only createSubnetwork(edges='all',subnetwork.name='top disease quartile connected') #handy way to exclude unconnected nodes! …from top quartile of ‘disease score’ top.quart top.quart)]ĬreateSubnetwork(top.nodes,subnetwork.name ='top disease quartile') In order to reflect your exploration back onto the network, let’s generate subnetworks…
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
